26 Nov. 2002 :
first release
18 Nov. 2004 :
slightly updated
R version is provided by Wolfram Stacklies. See http://rss.acs.unt.edu/Rdoc/library/pcaMethods/html/bpca.html .
Contents
Download
Zipped archive of MATLAB source files: BPCAFILL.zip(4kB)
Contents in the BPCAFILL.zip BPCAfill.m , BPCA_dostep.m , BPCA_initmodel.m , BPCA_reestimation.m , BPCA_plotM.m , missingNRMSE.m
Zipped archive of small sample data files: SAMPLE.zip(407kB)
Contents in the SAMPLE.zip sample999.dat , sampleorg.dat
Install
Place the source files (*.m) in arbitrary directory in MATLAB PATH.
Usage
Simplest usage
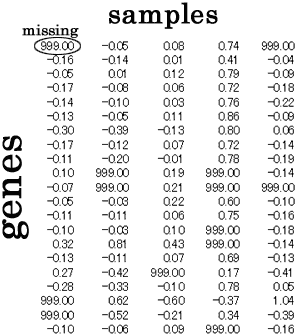
Prepare data matrix in which missing entries in matrix are
denoted by value 999.0.
BPCAfill estimates the missing entries using BPCA model
and returns complete matrix filled with the estimated values.
>> x999 = load('sample999.dat');
>> xfilled = BPCAfill( x999 );
epoch=10, dtau=1.25922
epoch=20, dtau=0.0970274
epoch=30, dtau=0.0349725
epoch=40, dtau=0.0150597
epoch=50, dtau=0.00761689
epoch=60, dtau=0.0034555
epoch=70, dtau=0.000254066
epoch=80, dtau=4.42614e-005 <-- when dtau < 1e-004, it is decided as converged.
>> save FilledExpression.dat -ascii xfilled
Please wait for the calculation to converge. It will take some minutes or hours till finish. (Example. The sample999.dat (757x50 entries with 797 missing) costs 10 minutes to fill using Pentium III 600MHz processer. )
Further analysis
-- Re-estimation
After the BPCAfill process, you can obtain a rough measure of the estimation acculacy prediction by using re-estimation process. BPCA_reestimation simulates 1% of artificial missing entries in the filled matrix, and estimates them again. The error on the re-estimation process will be a good prediction on the original error.
If the estimated NRMSE (normalized root mean squared error) value is near 1.00, the original fill might also not be reliable because of noise in the original expression data and/or other reasons.
>> [xfilled,M] = BPCAfill(sample999);
...
>> estimatedNRMSE = BPCA_reestimation(M);
rate =
0.0100
epoch=10, dtau=1.2374
...
epoch=80, dtau=5.04714e-005
>> estimatedNRMSE
estimatedNRMSE =
0.3664
>> xorg = load('sampleorg.dat');
>> trueNRMSE = missingNRMSE(x999, xorg, xfilled)
trueNRMSE =
0.4375
In the abobe, M and M2 are MATLAB structure variables which consist whole result of BPCA model estimation.
-- Show BPCA result
BPCA is a Bayesian variation of PCA, and it estimates covariance of the expression matrix as some principal axes. A profile of the result will be illustrated by BPCA_plotM.
>> [xfilled, M] = BPCAfill( x999 ); .... >> BPCA_plotM( M );